INTRODUCTION
MATERIALS AND METHODS
Experiment site and plant materials
Crop rotation and soil sampling
DNA extraction, PCR amplification, and sequencing
Microbiome taxonomic profiling and diversity indices analysis
Statistical analysis
RESULTS
Jack bean in crop rotation
Microbiome taxonomic profiling
Taxonomic composition analysis
Alpha and beta diversity analysis
DISCUSSION
INTRODUCTION
With agriculture facing the dual challenge of sustaining productivity and maintaining ecological balance, crop rotation has emerged as a highly effective approach to address these concerns (Shah et al., 2021; Shrestha et al., 2021). Crop rotation has numerous benefits, such as maintenance of soil health, and prevention of disease and pests (He et al., 2019; Yang et al., 2020). This practice also enhances crop productivity and economic efficiency (Acharya et al., 2020; Dias et al., 2015; Garbelini et al., 2022; Shah et al., 2021). By rotating crops, soil nutrients can be conserved in a balanced manner, and soil structure can be improved, leading to increased soil biodiversity and microbial activity (D’acunto et al., 2018).
The soil microbial community is a key component of soil health and plays a vital role in maintaining a healthy ecosystem (Chaparro et al., 2012; Sokol et al., 2022). This community is composed of a diverse range of microorganisms, including bacteria, fungi, protozoa, and other microscopic organisms. These organisms interact with each other and their environment, performing an array of ecological functions (Canarini et al., 2021; Romdhane et al., 2022). The interaction between plants and microorganisms affects the composition and function of microbial communities in the vicinity of plant roots, which are essential for maintaining plant health (Trivedi et al., 2020). Nutrient cycling is a crucial function within the soil microbial community, wherein microbes decompose organic matter and liberate essential nutrients such as nitrogen, phosphorus, and potassium, thereby making them accessible to plants (Yadav et al., 2021).
Legumes are integral to agriculture and ecosystems because of their unique ability to form symbiotic relationships with nitrogen- fixing bacteria, particularly Rhizobia (Andrews & Andrews, 2017). These interactions result in the formation of root nodules, allowing legumes to convert atmospheric nitrogen into a bioavailable form for plant growth (Ferguson et al., 2019). Therefore, the cultivation of legumes contributes to reduced dependence on synthetic nitrogen fertilizers and fosters sustainable agricultural practices. The use of legumes in crop rotation is gaining importance due to its positive effects on soil fertility and crop yield (Zhao et al., 2022). Several studies have explored the benefits of incorporating legumes such as soybean (Karlen et al., 1994; Sindelar et al., 2015), faba bean (Aschi et al., 2017; Jensen et al., 2010), and pea (Bailey et al., 2001; Matus et al., 1997) into crop rotation systems, demonstrating significant improvements in soil fertility and crop yield. Cultivation of legumes has also been reported to influence the composition, diversity, and function of soil microbial communities (Hartman et al., 2017; Wang et al., 2022). In the soybean rhizosphere, alterations in the microbial community structure were observed due to the increase of specific plant growth-promoting rhizobacteria (PGPR) genera (Sohn et al., 2021), and the cultivation of legumes has been reported to enhance soil microbial community activity and functional diversity, offering potential for ecological soil restoration (Li et al., 2012). Thus, understanding the dynamics of the soil microbiome in response to legume cultivation is crucial for developing agricultural systems that support increased productivity, biodiversity, and environmental sustainability.
The jack bean (Canavalia ensiformis), a member of the Fabaceae family, is commonly consumed by humans and used as animal feed. First cultivated in the West Indies and the adjacent mainland (Rana, 2017), it has recently been broadly distributed and cultivated in Asia (Akpapunam et al., 1997). Notably, jack bean exhibits tolerance to harsh soil conditions, including drought and extreme climates (Lynd & Ansman, 1989). This resilience extends to potential resistance against soil contamination. Owing to its rapid growth and extensive root system, jack bean demonstrates robust adaptability (Balliana et al., 2017). Similar to other legumes, jack bean can fix nitrogen and has shown high rates of growth in soils contaminated with multiple elements (Lynd & Ansman, 1989; Da Silva et al., 2018). Jack bean can also establish a mycorrhizal relationship with arbuscular mycorrhizal fungi, enabling the absorption and remediation of Zn- and Cu-contaminated soils (Andrade et al., 2009; 2010). However, compared to other legumes, little is known about the effect of jack bean cultivation on the soil microbial community.
This study seeks to explore the alterations of soil microbial community resulting from jack bean crop rotation. To achieve this, 16S rRNA gene sequencing was employed to analyze the microbial community composition under jack bean crop rotation.
MATERIALS AND METHODS
Experiment site and plant materials
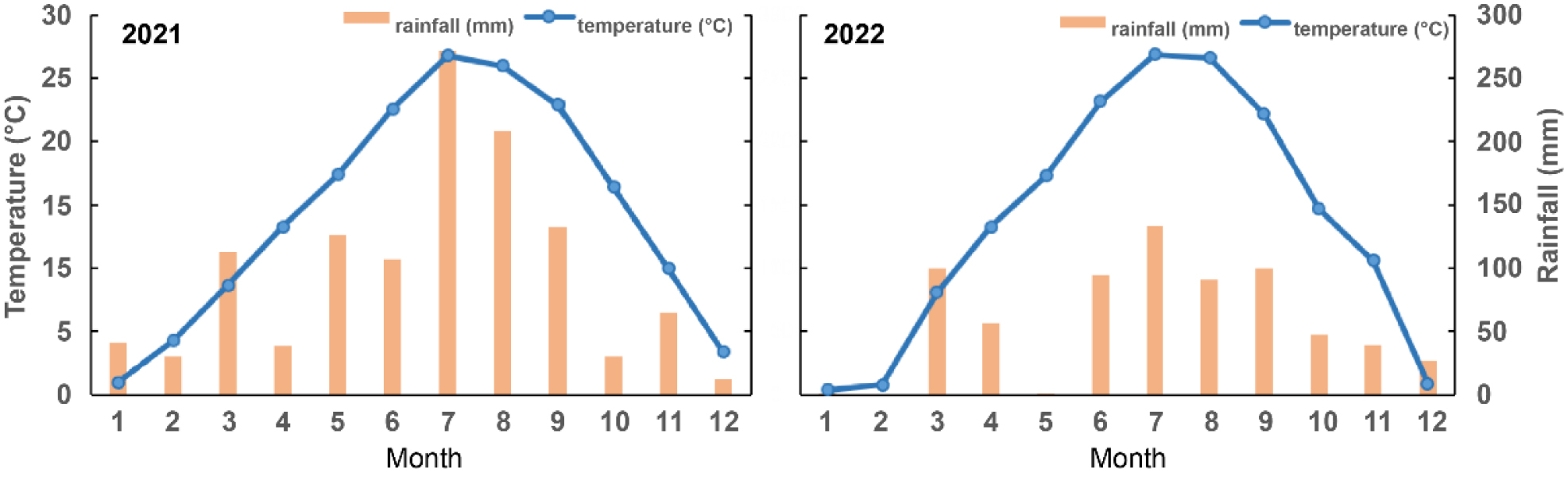
The experimental plot, with an area of 12 × 40 m2, was located at the Bioenergy Crop Research Institute, Muan, South Korea (34°58′ N, 126°27′ E), and the study was conducted 2011 and 2022. Prior to the field experiment, both physical and chemical properties of the soil were analyzed (Table 1). Additionally, average air temperature and precipitation throughout the crop cultivation period were recorded (Fig. 1). The onion and garlic cultivars used were ‘Mapsihwang,’ a mid-maturing cultivar, and ‘Daeseo,’ a southern-type cultivar predominantly cultivated in the southern regions of Korea. The jack bean used was an ‘SDG-4’ cultivar, which is extensively cultivated in the southern regions of Korea.
Table 1.
Physical and chemical properties of soil from the experimental area.
Crop rotation and soil sampling
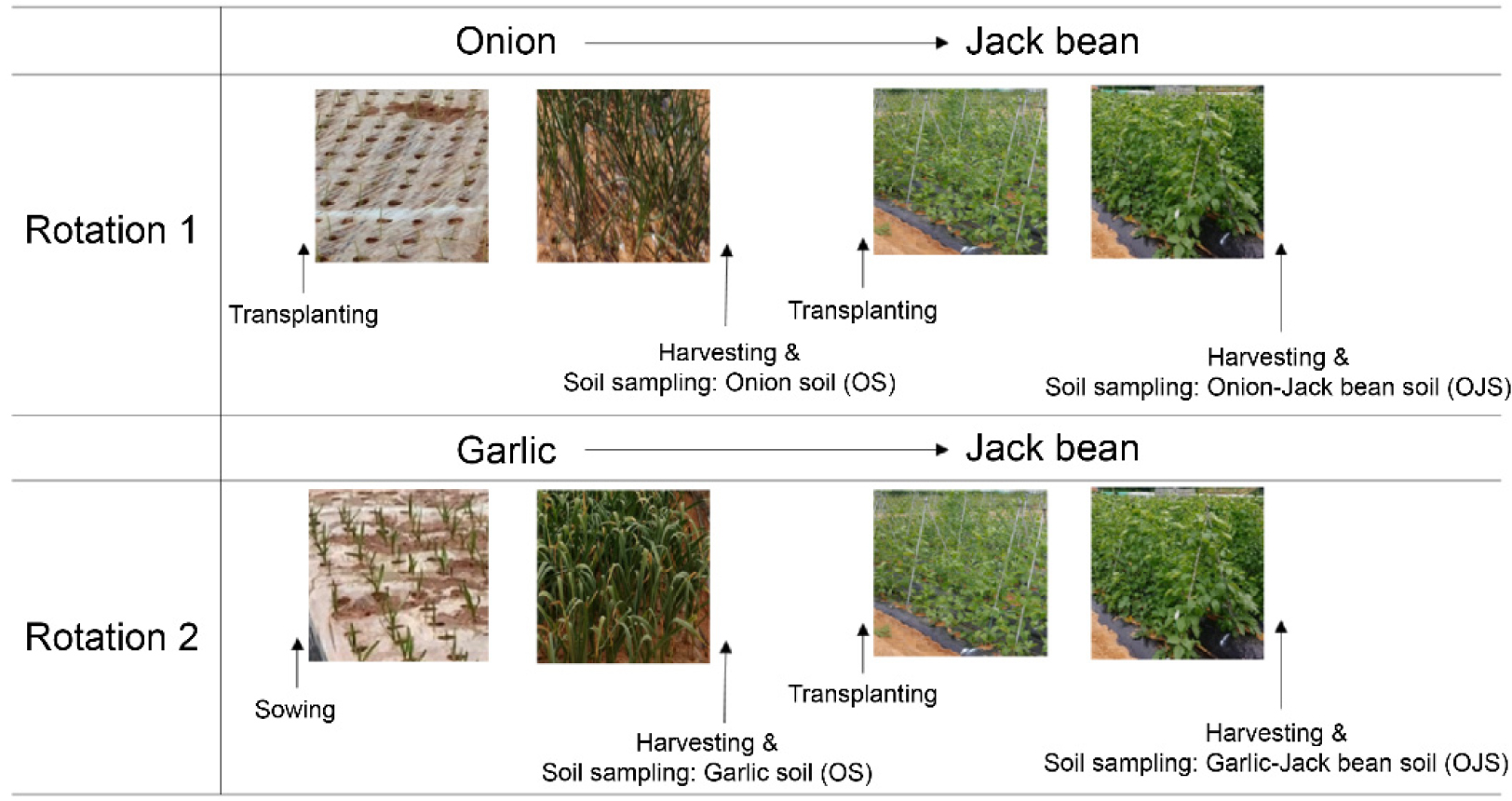
The experiment was conducted in soil that had only been used for cultivating onion and garlic. Crop rotation was conducted as follows: onion and garlic were cultivated from October 25, 2021, to May 19, 2022. Jack bean was sown in seed trays on May 2, 2022, and subsequently transplanted to the experimental plot on May 30, 2022, which had previously been used for onion and garlic cultivation. The crop was harvested on October 11, 2022. Before sowing or transplanting, fertilizers were applied as follows: onions received N-P-K at 80-77-58 kg/ha, garlic received N-P-K at 90-77-45 kg/ha, and jack bean received N-P-K at 57-90-68 kg/ha. After the wintering period, onions and garlic were top-dressed with N-K at 160-96 kg/ha and 160-83 kg/ha, respectively. The schematic representation of crop rotation and the timing of soil sampling are illustrated in Fig. 2. Jack bean straw was removed at harvest and not returned to the soil. The experimental plots for onion, garlic, onion-jack bean, and garlic-jack bean were replicated thrice, and rhizosphere soil samples were collected from nine sites in each plot. To isolate the rhizosphere soil, bulk soil was removed and the soil adjacent to the roots of the enlarged bulb of onion, garlic, and near the roots of jack bean was collected. The soil samples were immediately placed in ice packs and stored in a –30℃ deep freezer until further analysis.
DNA extraction, PCR amplification, and sequencing
Total DNA was extracted using the FastDNA Spin Kit (MP Biomedicals, California, USA) in accordance with the manufacturer’s instructions. The extracted DNA was used to perform PCR amplification targeting the V3 to V4 regions of the 16S rRNA gene, using fusion primers. For bacterial amplification, the fusion primers consisted of 341F (5’-TCGTCGGCAGCGTC- AGATGTGTATAAGAGACAG-CCTACGGGNGGCWGCAG-3’; underlining sequence indicates the target region primer) and 805R (5’-GTCTCGTGGGCTCGG-AGATGTGTATAAG AGACAG-GACTACHVGGGTATCTAATCC-3’) sequences. The fusion primers were constructed in the following order: P5(P7) graft binding, i5(i7) index, Nextera consensus, sequencing adaptor, and target region sequence. The PCR amplification was conducted with the following program: initial denaturation at 95℃ for 30 s, primer annealing at 55℃ for 30 s, and extension at 72℃ for 30 s, with a final elongation at 72℃ for 5 min. The PCR products were confirmed by analyzing with 1% agarose gel electrophoresis and visualized using a Gel Doc system (BioRad, Hercules, CA, USA). The purified PCR products were obtained with CleanPCR (CleanNA, Waddinxveen, Netherlands), and non-target products were removed. Equal amounts of purified products were combined and evaluated for quality and product size with a Bioanalyzer 2100 (Agilent, Palo Alto, CA, USA) using a DNA 7500 chip. The mixed amplicons were then sent for sequencing at CJ Bioscience, INC. (Seoul, Korea), using Illumina MiSeq sequencing system (Illumina, California, USA) following the manufacturer’s protocol.
Microbiome taxonomic profiling and diversity indices analysis
Processing raw reads started with quality check (QC) and filtering of low quality (<Q25) reads by Trimmomatic ver. 0.32 (Bolger et al., 2014). After QC pass, paired-end sequence data were merged together using fastq_mergepairs command of VSEARCH version 2.13.4 (Rognes et al., 2016) with default parameters. Primers were then trimmed with the alignment algorithm of Myers & Miller (Myers & Miller, 1988) at a similarity cut off of 0.8. Non-specific amplicons that did not encode 16S rRNA were detected by nhmmer (Wheeler et al., 2013) in HMMER software package ver. 3.2.1 with hmm profiles. Unique reads were extracted and redundant reads were clustered with the unique reads by derep_fullength command of VSEARCH. The EzBioCloud 16S rRNA database was used for taxonomic assignment using usearch_global command of VSEARCH followed by more precise pairwise alignment.
Chimeric reads were filtered on reads with <97% similarity by reference-based chimeric detection using UCHIME algorithm and the non-chimeric 16S rRNA database from EzBioCloud. After chimeric filtering, reads that are not identified to the species level (with <97% similarity) in the EzBioCloud database were compiled and cluster_fast command was used to perform de-novo clustering to generate additional operational taxonomic units (OTUs). Finally, OTUs with single reads (singletons) were omitted from further analysis.
The secondary analysis, which includes diversity calculation, was conducted by in-house programs of CJ Bioscience, Inc (Seoul, South Korea). The alpha diversity indices such as ACE (Abundance-based Coverage Eistmator) (Chao & Lee, 1992), Chao1 (Chao, 1987), Jackknife (Burnham & Overton, 1979), Shannon (Magurran, 2013), NPShannon (Chao & Shen, 2003), and Phylogenetic diversity (Faith, 1992) were estimated. To visualize the sample differences, beta diversity distances were calculated by principal coordinate analysis (PCoA) based on Jensen-Shannon algorithms (Lin, 1991).
Statistical analysis
The results are reported as the mean ± standard deviation of triplicate experiments. One-way analysis of variance (ANOVA) with Duncan’s multiple range test (p ≤ 0.05) was performed with R-Studio ver. 1.3.1073.
RESULTS
Jack bean in crop rotation
The physical and chemical properties of the soil are presented in Table 1. The soil texture was classified as loam. The soil pH was determined to be slightly acidic at 5.5, and the electrical conductivity (EC) was 0.59 dS/m, indicating a low salinity level. The chemical properties of the soil showed significant changes across the different stages of crop cultivation. Organic matter and available P₂O₅ levels were low before the experiment but increased after the cultivation of onion and garlic. These values were particularly high after the cultivation of jack bean compared to the onion and garlic cultivation soils. Additionally, exchangeable potassium (Ex. K), calcium (Ex. Ca), magnesium (Ex. Mg), and sodium (Ex. Na) were higher in the soils after the cultivation of onion, garlic, and jack bean compared to the pre-experiment soil. However, there were no significant differences among the crops in terms of these exchangeable cations. During the jack bean growth season (30 May–11 October), the total precipitation amounted to 468.0 mm (Fig. 1). The temperature peaked in July following transplantation and subsequently declined. Approximately 60 days post-transplantation, young seedpods of jack bean were observed, and these were harvested before sowing garlic and onion.
Microbiome taxonomic profiling
A total of 448,311 pre-filtered reads were obtained from 12 soil samples, representing four experimental plots—onion soil (OS), garlic soil (GS), onion-jack bean soil (OJS), and garlic-jack bean soil (GJS)—with three replicates each.. After the removal of low-quality, non-target, and chimeric amplicons, a total of 402,845 valid reads were obtained. In the 12 soil samples, overall, 323,399 reads were identified at the species level using reference databases, and the number of unique species identified amounted to 28,853 reads. The read lengths ranged from 110 bp to 467 bp, with an average of 413 bp (Table 2).
Table 2.
Sequencing and taxonomic identification metrics from reads.
| MTP | OS | GS | OJS | GJS | |
| Total reads after pre-filter | 36814.0 ± 2342.6 | 43749.3 ± 13810.4 | 33919.3 ± 2461.0 | 34954.3 ± 2063.1 | |
| Total valid reads | 33439.0 ± 2153.8 | 39537.3 ± 12933.3 | 30193.0 ± 2182.7 | 31112.3 ± 1776.1 | |
| Low quality amplicons | 2678.0 ± 109.7 | 2997.3 ± 582.4 | 3087.7 ± 368.0 | 3272.7 ± 338.1 | |
| Non-target amplicons | 251.3 ± 99.9 | 494.7 ± 122.5 | 59.7 ± 1.5 | 41.7 ± 4.2 | |
| Chimeric amplicons | 445.7 ± 97.2 | 720.0 ± 265.1 | 579.0 ± 97.9 | 527.7 ± 72.0 | |
| No. of reads identified at the species level | 27278.7 ± 2161.0 | 31451.0 ± 10200.8 | 15804.7 ± 1709.1 | 24611.7 ± 1495.9 | |
| No. of species found | 2708.7 ± 96.0 | 2835.3 ± 308.0 | 2083.7 ± 58.4 | 1990.0 ± 76.2 | |
| Read lengths | Min | 120 | 120 | 110 | 120 |
| Max | 448 | 467 | 456 | 448 | |
| Average | 414.8 ± 0.4 | 414.8 ± 0.1 | 410.7 ± 0.3 | 410.8 ± 0.2 | |
Taxonomic composition analysis
At the phylum level, excluding those with an average abundance under 1% (Others), 10 phyla were identified in OS and GS, while nine phyla were found in OJS and GJS (Fig. 3). The dominance of Proteobacteria, Actinobacteria, Acidobacteria, and Bacteroidetes was consistent across all soil types. Proteobacteria stood dominant across all soil types, with relative abundances of 42.51%, 42.07%, 47.72%, and 48.7% in OS, GS, OJS, and GJS, respectively, and Actinobacteria was the second most dominant in all soil types, with abundances of 16.32%, 14.48%, 22.3%, and 20.16%, respectively. It is noteworthy that Firmicutes, which recorded 4.85% in OS and 5.48% in GS, were conspicuously absent in OJS and GJS. In OJS and GJS, there was a discernible increase in the presence of Proteobacteria, Actinobacteria, and Saccharibacteria, whereas phyla such as Verrucomicrobia, Chloroflexi, Gemmatimonadetes, and Bacteroidetes declined. Furthermore, Alphaproteobacteria was the only subgroup that showed an increase when the prevalent Proteobacteria in OJS and GJS was considered (Fig. 4).
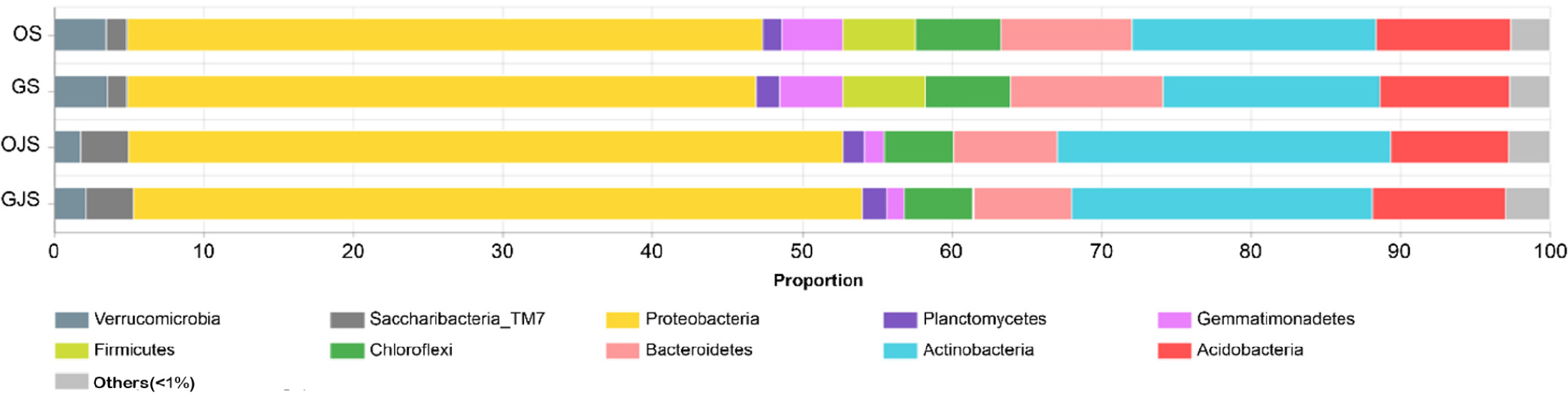
Fig. 3.
Taxonomic composition of soil microbial communities at the phylum level based on soil samples collected after cultivating onion, garlic, and jack bean. The bars represent the average relative abundance from triplicate samples, and labels indicate taxonomic units with an average relative abundance >1%. OS, onion soil; GS, garlic soil; OJS, onion-jack bean soil; GJS, garlic-jack bean soil. Across all soil types, Proteobacteria and Actinobacteria were the dominant phyla. Notably, both phyla showed higher relative abundances following jack bean cultivation (OJS and GJS).
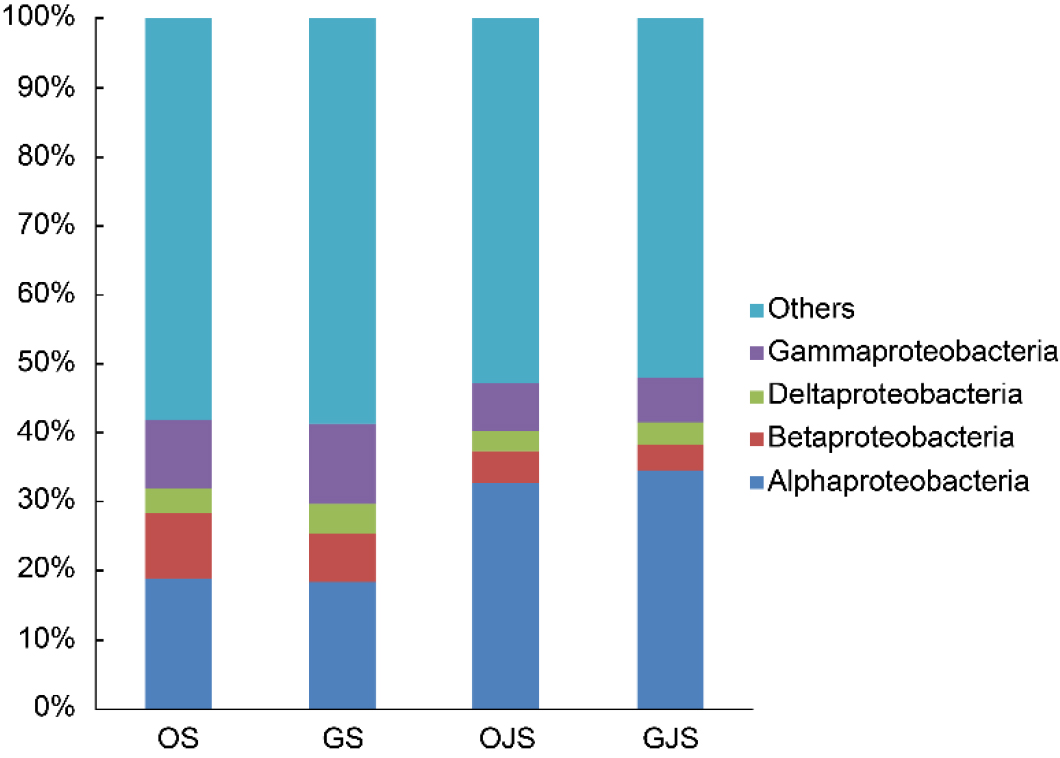
Fig. 4.
Taxonomic composition of the soil microbial communities at the class level based on soil samples collected after cultivating onion, garlic, and jack bean. The bars represent the average relative abundance from triplicate samples. OS, onion soil; GS, garlic soil; OJS, onion-jack bean soil; GJS, garlic-jack bean soil. Within Proteobacteria, the proportion of Alphaproteobacteria, a class that includes many functionally important genera, was higher in the jack bean-cultivated soils (OJS and GJS) than in the pre-cultivated soils (OS and GS).
At the genus level, a total of 31 genera were identified in the four soil types. Excluding ETC (under 1% on average), 14 genera were observed in OS and GS, while 19 genera in OJS, and 20 genera in GJS (Fig. 5). OS and GS predominantly comprised genera such as Sphingomonas (5.59% and 4.91%, respectively) and Arthrobacter (5.82% and 4.41% in Garlic). Although relative abundance varied depending on the previous crop, Sphingomonas (4.85% and 4.48%, respectively) and Pseudolabrys (3.82% and 4.86%, respectively) exhibited a dominant tendency in OJS and GJS. Microorganisms that were not detected in onion and garlic soils were found in jack bean soil, including Bauldia, Devosia, Pseudolabrys, Rhizobium, Actinoplanes, and Bradyrhizobium. In contrast, some microorganisms that were detected in onion and garlic soils were not found in jack bean soil. Lysobacter and Nitrosospira were found in OS, and Chryseolinea and Aquicella were found in GS, but were not found in OJS and GJS.
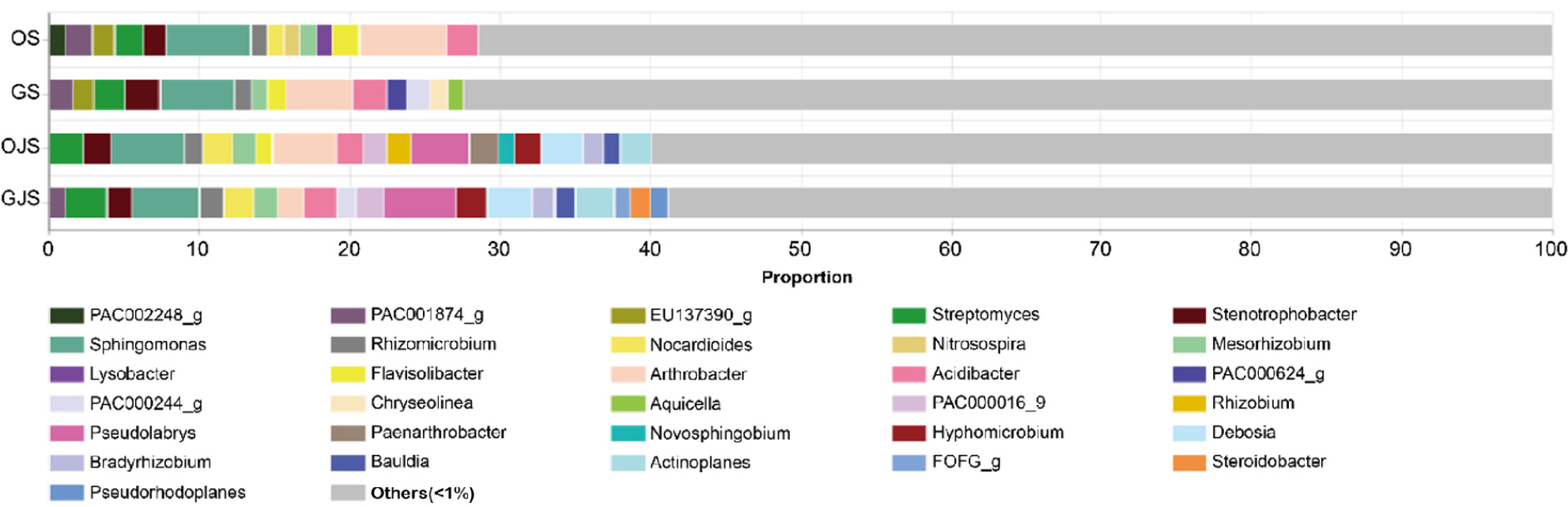
Fig. 5.
Taxonomic composition of soil microbial communities at the genus level, based on soil samples collected after cultivating onion, garlic, and jack bean. The bars represent the average relative abundance of triplicate samples, and labels indicate the taxonomic units with an average relative abundance >1%. OS, onion soil; GS, garlic soil; OJS, onion-jack bean soil; GJS, garlic-jack bean soil. Soils follwing jack bean cultivation (OJS and GJS) included key functional genera such as Pseudolabrys, Bradyrhizobium, Devosia, Bauldia, and Rhizobium, which were not detected in the soils prior to jack bean cultivation (OS and GS).
Alpha and beta diversity analysis
To analyze the richness and diversity of the soil microbial community following jack bean cultivation, species richness (the number of species in MTP) was assessed using ACE, Chao1, and Jackknife indices. Additionally, diversity indices (species diversity or evenness in MTP) were calculated using Shannon, NPShannon, and Phylogenetic diversity. All the values (higher values indicate higher diversity) were higher in OS and GS than in OJS and GJS (Table 3).
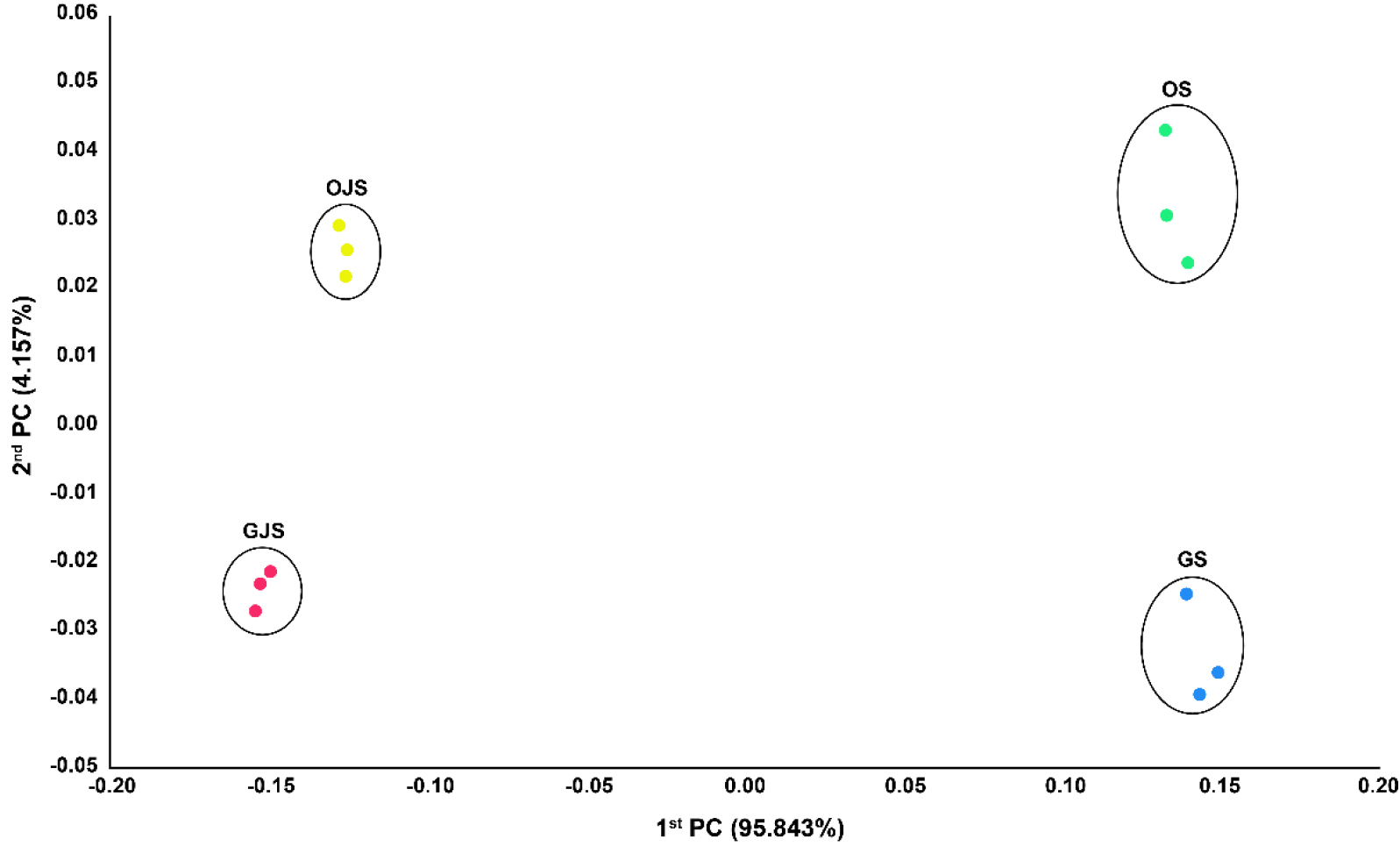
Changes in soil microbial community in each soil were analyzed using principal coordinate analysis (PCoA) based on Jensen-Shannon divergence distance (Fig. 6). The PCoA results demonstrated that the microbial community composition was different before and after jack bean cultivation, as represented by the first principal coordinate (PC1, 95.843% of variance). The second PC (4.157% of total variance) showed clear clustering based on the type of crops, indicating that the soil microbial community was influenced not only by jack bean cultivation but also by the previous crop as well as various seasonal and environmental conditions throughout the growing season.
DISCUSSION
This study investigated the changes in soil microbial communities induced by the cultivation of jack bean, which is primarily grown in winter following the cultivation of onion and garlic in the southern region of Korea. Through 16S rRNA gene sequencing, shifts in the soil microbial community post-jack bean cultivation were identified, along with the emergence of microbes typically found in other legumes. These findings establish a foundation for implementing diversified crop rotations with jack bean and provide essential data for understanding its nitrogen fixation mechanisms and agricultural benefits.
Alpha diversity analyses consistently demonstrated comparable patterns both before and after jack bean cultivation. Indices such as ACE, Chao1, Jackknife, Shannon, NPShannon, and Phylogenetic diversity were higher in soils previously cultivated with onions and garlic compared to post-jack bean cultivation. This pattern led us to hypothesize that the reduction in species richness and diversity within the soil microbial community following jack bean cultivation was due to the dominance of specific microorganisms. In a study analyzing the microbial community across different growth stages of soybean, while a higher number of genera were detected during its later growth stages (R5 to R7) compared to the pre-planting bulk soil, the diversity indices were lower (Sohn et al., 2021). This observation is consistent with findings from post-jack bean cultivation soils (OJS and GJS), in which more genera were detected compared to pre-jack bean cultivation soils (OS and GS), despite having lower diversity indices values. Diversity indices that reflect species diversity or evenness, such as Shannon, NPShannon, and Phylogenetic diversity, were higher in OS and GS than in OJS and GJS. The higher number of genera results in lower diversity indices due to lower overall evenness; specific genera may rapidly increase under certain environmental conditions, causing the relative proportions of other genera to decrease.
This implies that although the richness of genera increases, the community becomes less balanced, reflecting changes in the dominance and distribution of microbial populations According to Ma (2005) and Hillebrand et al. (2008), decreased evenness significantly contributes to the decline in overall diversity indices. PCoA analysis demonstrated that the bacterial communities post-jack bean cultivation formed distinct clusters. Within the PCoA space, distinct clusters were observed for both post- (OJS and GJS) and pre-jack bean cultivation soils (OS and GS), indicating that jack bean cultivation affects the composition of the soil microbial community.
In all the examined soils, Proteobacteria was the most predominant phylum, followed by Actinobacteria, Acidobacteria, and Bacteroidetes. Significantly, both Proteobacteria and Actinobacteria demonstrated a notably higher distribution ratio post-jack bean cultivation compared to pre-cultivation periods, aligning with previous reports on the rhizosphere soil microbial communities of soybean, pea, and alfalfa (Mendes et al., 2014; Xiao et al., 2017; Qi et al., 2023; Chaudhari et al., 2020). As a phylum, Proteobacteria, a phylum of Gram-negative bacteria, is commonly found in soil environments and is associated with a wide range of functions related to the cycling of carbon, nitrogen, and sulfur (Spain et al., 2009). The observed increase in Proteobacteria distribution was primarily due to the increase in Alphaproteobacteria (Fig. 4), which are known for their roles in degradation of inorganic compounds and facilitating nitrogen fixation (Liu & Liu, 2013). Actinobacteria are also known to play a significant role in ecosystems (Hill et al., 2011). They produce a variety of extracellular hydrolytic enzymes that decompose other organic compounds (Eisenlord & Zak, 2010) and can play a crucial role in nitrogen cycling (Zhang et al., 2019). Firmicutes, found in the post-onion and garlic cultivation soil, were absent in the post- jack bean cultivation, and the distribution ratios of Chloroflexi and Gemmatimonadetes also decreased. This phenomenon is hypothesized to have occurred as a result of other microbial communities becoming dominant. Firmicutes, Chloroflexi, and Gemmatimonadetes are known as oligotrophic communities that thrive in nutrient-depleted and disturbed environments (Csecserits et al., 2011; Davis et al., 2011; Koyama et al., 2014; Lladó et al., 2017). This result is associated with the higher organic matter observed after jack bean cultivation compared to that after onion and garlic harvests (Table 1).
The microbial dynamics within soils cultivated for onion, garlic, and jack bean reveal complex interactions with potential implications for nutrient cycling and plant growth promotion. Sphingomonas, found to be abundant in onion, garlic, and jack bean soils, are recognized for their production of indole-3-acetic acid (Tsavkelova et al., 2007) and their involvement in the degradation or transformation of toxic heavy metals and other environmental pollutants (Mahbub et al., 2016). Legume plants engage in symbiotic relationships with Rhizobium and other microbes. These symbiotic relationships convert nitrogen into a form usable by the plant. Our study notably observed an abundance of these microbes in the soil following the cultivation of jack bean. Pseudolabrys, Bradyrhizobium, and Devosia are known to play substantial roles in nitrogen cycling (Yu et al., 2022) and have been proven to be involved in nitrogen fixation and nitrification (Barbieri et al., 2010; Gohar et al., 2020; Rivas et al., 2002). Rhizobium is the most important plant growth- promoting rhizobacteria (PGPR) (Miransari, 2016). Rhizobium live in the soil or within legume root nodules and have the ability to fix atmospheric nitrogen, thereby providing plants with nitrogen (Ulzen et al., 2016). In addition to nitrogen fixation, Rhizobium also produces amino acids, siderophores, and auxins (Boiero et al., 2007), as well as indole acetic acid (IAA) and gibberellic acid (GA) (Egamberdieva et al., 2017), which collectively contribute to enhancing root growth (Purwaningsih et al., 2021). Actinoplanes, known for producing phytohormones, siderophores, and facilitating phosphate solubilization, also play a crucial role in the soil microbial community (Menendez & Carro, 2019). The increased distribution rates of Proteobacteria and Alphaproteobacteria following jack bean cultivation can be attributed to the rise in populations of Sphingomonas, Pseudolabrys, Bradyrhizobium, Devosia, Buldia, and Rhizobium, which belong to these phylum and class. These microorganisms are known for their positive impact on soil health, indicating their crucial role in enhancing soil quality post-jack bean cultivation. Previous studies have also reported that genera enriched follwing jack bean cultivation, particularly Bradyrhizobium, Rhizobium, and Sphingomonas are similarly dominant in the rhizospheres of other legumes such as soybean and pea (Sohn et al., 2021; Chaudhari et al., 2020). Therefore, the changes in the bacterial community due to jack bean cultivation suggest the formation of a unique bacterial community associated with jack bean cultivation.
Intensive agricultural practices such as continuous monocropping and massive chemical input have been shown to significantly alter belowground microbial composition and function, often resulting in negative feedback on soil diseases, crop growth, and production (Ehrenfeld et al., 2005; Mazzola & Manici, 2012; Wang et al., 2022). Such practices can lead to unhealthy soil conditions due to imbalanced nutrient depletion (Ehrenfeld et al., 2005), diminished activity of beneficial microbes (Wang et al., 2019), disrupted soil microbial communities (Li et al., 2020; Liu et al., 2014), and simplified microbial networks (Lu et al., 2013). These unfavorable outcomes often result in significant crop yield losses.
To counter these problems, diverse cropping systems such as crop rotation and intercropping are being implemented. Crop rotation can disrupt the cycle of weeds and diseases (Andam et al., 2016), while also enhancing soil fertility through increased soil microbial activity and higher organic matter content (Bullock, 1992). Particularly, the use of legumes as rotational crops is encouraged to maintain crop yield and preserve soil quality (Luce et al., 2015; Yusuf et al., 2009). The aim of this study was to anticipate the comprehensive effects of crop rotation by gaining an in-depth understanding of microbial diversity and the role of key genera following the cultivation of jack bean.
However, further research is needed to understand specific interactions between these bacterial communities and other factors influencing soil health, crop growth, and the ecosystem. For example, studies comparing fallow land with jack bean- cultivated soil and the effects of returning jack bean straw to the soil are necessary. This study has certain limitations, as the experiment was conducted with only three biological replicates per treatment and sampling was performed at a single post-harvest timepoint. It is important to investigate the long-term effects of jack bean cultivation on soil microbial communities in order to better understand the sustainability of this cropping practice. Future studies should incorporate time-series sampling and increased replication to more precisely capture the temporal and spatial dynamics of microbial community changes over time. Furthermore, in this study, the functional interpretation of microbial communities was based solely on taxonomic classification. Such an approach may not fully reflect the actual metabolic or ecological functions of the microbes. To address this limitation, future studies should incorporate gene-based functional validation methods or predictive tools such as PICRUSt or Tax4Fun to more accurately evaluate the functional potential of microbial communities associated with jack bean cultivation.