INTRODUCTION
MATERIAL AND METHODS
SNDH population and QTL mapping
Infection assays on the SNDH population
Prediction of potential genes using bioinformatics tools
RESULTS
Phenotype evaluation for rice blast resistance in the SNDH population
QTL analysis for rice blast resistance in the SNDH population
Candidate genes for rice blast resistance within major QTLs
Selection of the target gene for rice blast resistance
Phylogenetic and protein-protein interaction analysis of the target gene Osrbrq2
DISCUSSION
CONCLUSIONS
INTRODUCTION
Rice (Oryza sativa L.) is a primary food source for more than half of the global population, making sustained productivity essential for global food security (Bandumula, 2018). Among the biotic stresses that threaten rice cultivation, blast disease—caused by the ascomycete fungus Magnaporthe oryzae—remains the most destructive (Liu et al., 2024). The pathogen can infect all aerial parts of the rice plant, often resulting in severe yield losses under favorable environmental conditions (Zhang et al., 2024). Although the deployment of resistant cultivars is central to effective blast management (Sharma et al., 2012). However, the durability of resistance is frequently undermined by the rapid evolution and regional diversification of pathogen populations (Pedrozo et al., 2025).
Quantitative trait locus (QTL) analysis has become a key tool for dissecting the genetic basis of complex traits, including disease resistance in rice (Zhao et al., 2024; Sharma et al., 2021). By linking phenotypic variation to specific genomic regions, QTL mapping facilitates the identification of loci and candidate genes that contribute to quantitative and often polygenic resistance. This approach has been widely applied in rice blast research to identify stable, environmentally robust loci suitable for breeding programs (Liu et al., 2021). Recent advances in high-density genotyping, improved mapping populations, and integrative genomic analyses have further enhanced the precision and utility of QTL-based studies in uncovering resistance- associated genes (Liu et al., 2021; Sallaud et al., 2003). Consequently, QTL analysis now serves as a powerful complement to traditional phenotypic screening, enabling breeders to develop blast-resistant rice cultivars more efficiently and with greater genetic insight.
Recently, the race distribution and pathogenic diversity of the rice blast fungus in Korea were characterized using regional isolates collected from 2020 to 2022. From this collection, a set of 28 representative isolates belonging to different pathotypes was selected based on their viability, high virulence, predominant race types, and distinct reaction patterns to known blast resistance genes identified in a previous study (Zhao et al. 2026, in press).
In this study, QTL analysis was conducted using these 28 representative Korean isolates and a doubled haploid population derived from a cross between the blast-resistant cultivar ‘Samgang’ and the susceptible cultivar ‘Nagdong’. The primary objective was to identify key QTLs and associated candidate genes linked to rice blast resistance. The findings provide valuable genetic resources that can support breeding programs aimed at developing resilient, broad-spectrum rice cultivars with enhanced blast resistance tailored to Korean rice-growing environments.
MATERIAL AND METHODS
SNDH population and QTL mapping
A total of 101 SNDH lines were developed through anther culture from the F1 cross between the blast-resistant Indica cultivar ‘Samgang’ and the blast-susceptible Japonica cultivar ‘Nagdong’ at the Plant Molecular Breeding Laboratory, Kyungpook National University, Republic of Korea (Park et al., 2020). This population was used to identify QTLs associated with rice blast resistance. Among 850 SSR markers initially screened, 163 polymorphic and codominant markers exhibiting clear amplification patterns were selected for the construction of the genetic linkage map. Linkage analysis was conducted using MAP Manager QTXb2.0, and marker distances were calculated with the Kosambi mapping function (Manly & Olson, 1999). Composite interval mapping was performed using WinQTL Cartographer 2.5 to detect significant QTLs, employing a logarithm of odds (LOD) threshold of 3.0 (Zeng, 1994). The proportion of phenotypic variation explained by each QTL was expressed as the R2 value. Identified loci were named according to the guidelines proposed by McCough & Doerge (1995).
Infection assays on the SNDH population
Infection assays were conducted on the SNDH population using a conidial mixture prepared from 28 representative M. oryzae isolates selected between 2020 and 2022 in a previous study (Zhao et al. 2026, in press). Each isolate was initially cultured on potato dextrose agar for 7 days and subsequently subcultured on rice bran agar for an additional 10 days (Fig. S1). Conidiation was induced by exposing the cultures to UV light for 3 days. The resulting conidia were harvested, suspended in sterile water containing Tween 20 (250 ppm; CAS: 9005-64-5), and adjusted to a concentration of 1 × 105 conidia/mL. Seeds from each SNDH line were sown in 72-well seedling trays filled with sterilized nursery soil and maintained in greenhouse conditions until the four-leaf stage, at which point plants were used for inoculation. The mixed conidial suspension was sprayed onto 101 SNDH lines along with the parental varieties ‘Samgang’ and ‘Nagdong’. Five seedlings per line were inoculated, and each assay was performed in triplicate. Following inoculation, plants were placed in a dew chamber at 26°C for 24 h in darkness, then transferred to a greenhouse with a 16-h light/8-h dark photoperiod. Disease severity was assessed seven days after inoculation using a 1–5 scale, where 1 = uniform dark-brown pinpoint lesions lacking visible centers and 5 = large eyespot lesions (~5 mm) (Valent et al., 1991). Representative infected leaves were photographed for documentation (Table S1).
Prediction of potential genes using bioinformatics tools
To identify potential candidate genes associated with rice blast resistance, bioinformatic analyses were performed on the major QTL regions detected in the SNDH population. The genomic intervals flanked by significant SSR markers were first examined using the Rice Annotation Project Database (RAP- DB) (Sakai et al., 2013) and the RiceXPro expression database (Sato et al., 2013) to obtain positional information and expression profiles of genes located within or near the identified marker regions. Predicted open reading frames were annotated based on their putative biological functions, with particular attention to genes previously implicated in plant defense, including receptor- like kinases, transcription factors, and pathogenesis-related proteins. Gene ontology (GO) enrichment analysis was conducted using the agriGO v2.0 platform (Job ID: 599983822.1) to evaluate the biological significance of the candidate genes, categorizing them into biological processes, molecular functions, and cellular components (Du et al., 2010). Sequence similarity analyses were carried out using the National Center for Biotechnology Information database, and a phylogenetic tree of the target gene was constructed using MEGA12. Furthermore, protein–protein interaction networks for the target gene were examined using the STRING database (Version 12.0) (Szklarczyk et al., 2019).
RESULTS
Phenotype evaluation for rice blast resistance in the SNDH population
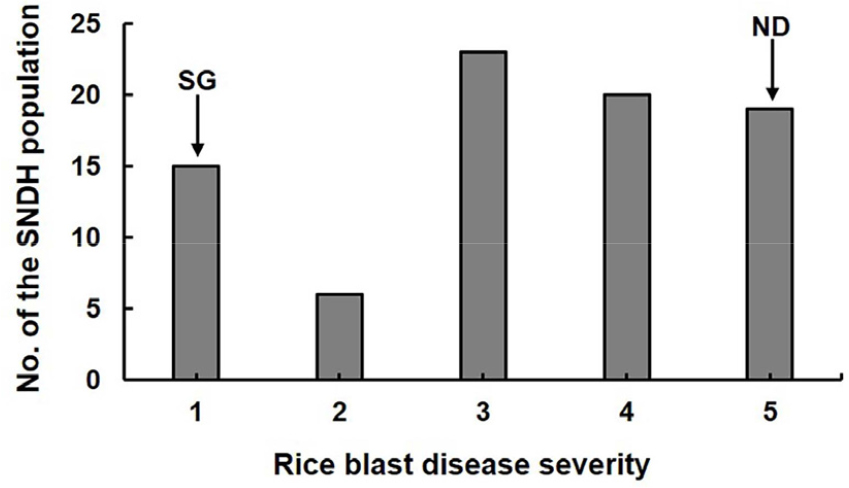
For QTL analysis, disease severity in the SNDH population was evaluated seven days after inoculation with a mixture of 28 representative M. oryzae isolates. Disease response was scored on a numerical scale from 1 to 5. Severity scores among the SNDH lines ranged from 1 to 5, with an average of 3.27, reflecting substantial phenotypic variation from moderately resistant to highly susceptible. These data were used to construct a frequency distribution illustrating resistance pattern within the population (Fig. 1). The parental cultivars displayed the expected phenotypes: the blast-resistant Indica cultivar ‘Samgang’ exhibited minimal symptoms with a score of 1, whereas the susceptible Japonica cultivar ‘Nagdong’ showed severe disease symptoms with a score of 5 (Table S1).
QTL analysis for rice blast resistance in the SNDH population
QTL analysis was performed using the rice blast disease severity data from the SNDH population to identify genomic regions associated with resistance. Two major QTLs were detected with significant associations to rice blast resistance. The first QTL, designated qrbr2, was located within the RM318–s2038 marker interval on chromosome 2 and explained 44% of the phenotypic variation. The second QTL, qrbr6, was identified within the s6015–s6018 marker interval on chromosome 6 and accounted for 43% of the variation (Fig. 2). The LOD values for these loci were 4.13 for qrbr2 and 5.15 for qrbr6, confirming strong statistical significance (Table 1, Fig. 3A). In both cases, the resistant alleles originated from the blast-resistant parent cultivar ‘Samgang,’ highlighting its substantial contribution to resistance in the SNDH population. The identification of qrbr2 and qrbr6 provides valuable targets for marker-assisted selection and future functional characterization.
Table 1.
Significant QTLs associated with rice blast disease resistance were identified in the SNDH population.
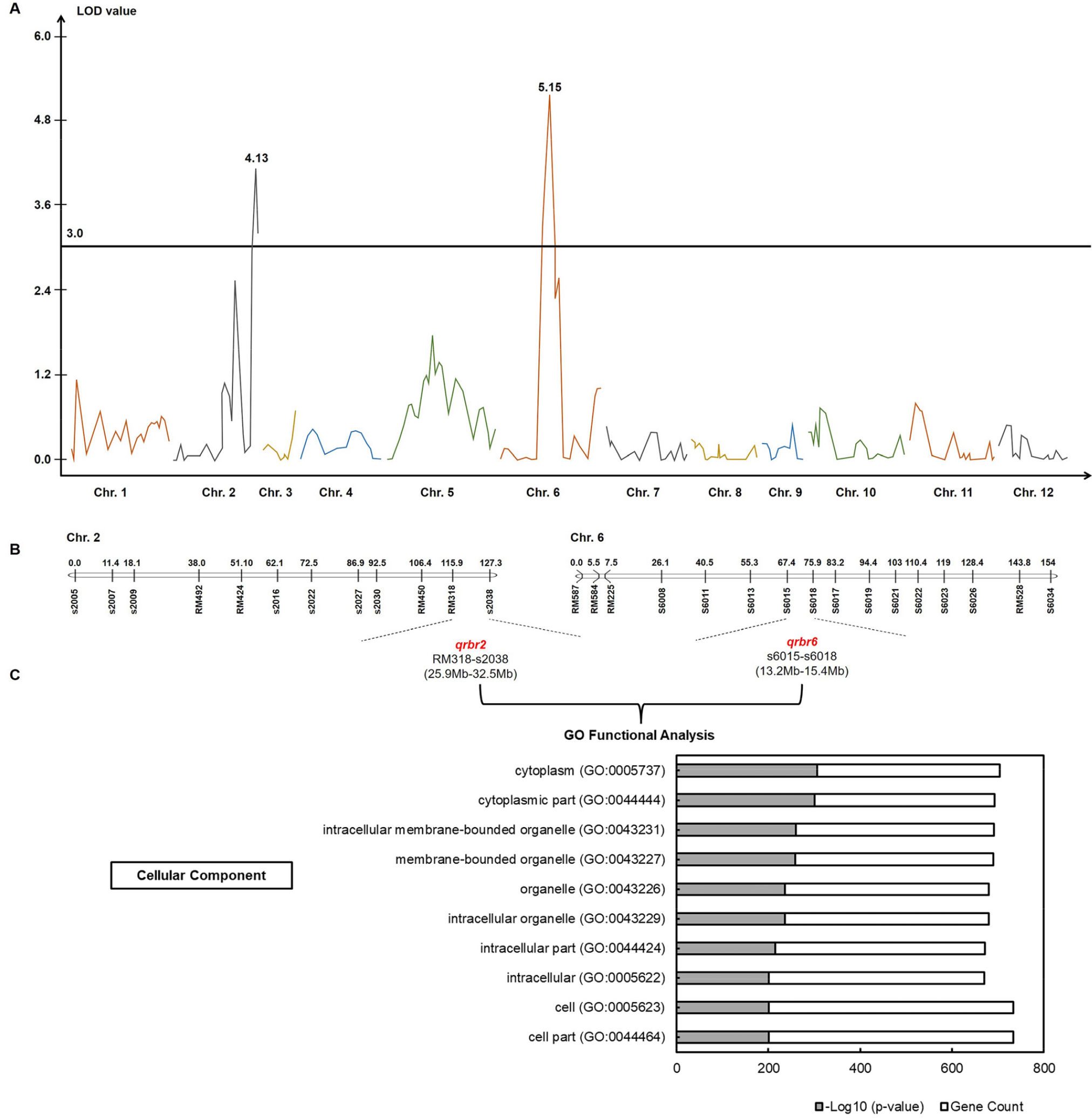
Fig. 3.
QTL analysis for rice blast resistance and GO functional analysis of genes within target QTLs. (A) Two QTLs, qrbr2 and qrbr6, were identified on chromosomes 2 and 6, respectively. (B) Target marker intervals, RM318-s2038 and s6015-s6018, were detected on chromosomes 2 and 6, respectively. (C) Top ten GO terms associated with Cellular Component for genes within the QTL regions.
Candidate genes for rice blast resistance within major QTLs
To identify candidate genes potentially associated with rice blast resistance, the genomic regions corresponding to the two major QTLs—RM318–s2038 on chromosome 2 (qrbr2) and s6015–s6018 on chromosome 6 (qrbr6)—were examined (Fig. 3B). Gene annotation and expression information were retrieved from the RiceXPro and RAP-DB databases. A total of 774 genes were screened, with 701 genes located within the qrbr2 and 73 genes within the qrbr6 region (Table S2). Functional insights were obtained through GO analysis, which revealed several significantly enriched GO terms. Among these terms, the ten most abundant terms belonged to the Cellular Component category and included cytoplasm (GO:0005737), cytoplasmic part (GO:0044444), intracellular membrane-bounded organelle (GO:0043231), membrane-bounded organelle (GO:0043227), intracellular organelle (GO:0043229), organelle (GO:0043226), intracellular part (GO:0044424), intracellular (GO:0005622), cell (GO:0005623), and cell part (GO:0044464) (Fig. 3C).
Selection of the target gene for rice blast resistance
Annotation-based classification of the 774 candidate genes revealed a broad distribution of functional groups. The five largest categories included conserved hypothetical proteins (160 genes), hypothetical proteins (24 genes), nonprotein-coding genes (15 genes), zinc finger RING-type domain-containing proteins (11 genes), and protein prenyltransferase domain- containing proteins (10 genes) (Table S2). Among these, the zinc finger RING-type domain-containing proteins were prioritized for further investigation, as this class of E3 ubiquitin ligases is widely recognized for its involvement in plant defense regulation and intracellular signaling pathways. Their roles in recognizing and modulating defense-related transcription factors have been documented in several studies on rice blast resistance. Of the 11 genes within this category, we selected Os02g0674700, designated Osrbrq2, as the primary target for downstream functional analysis because of its proximity to the peak of the QTL locus qrbr2.
Phylogenetic and protein-protein interaction analysis of the target gene Osrbrq2
QTL mapping identified the candidate gene Osrbrq2, associated with rice blast resistance, within the qrbr2 locus (Fig. 4A). Phylogenetic analysis clarified the genetic relationships of Osrbrq2 with homologs in Triticum dicoccoides, Triticum aestivum, Hordeum vulgare subsp. vulgare, and Oryza glaberrima (Fig. 4B). Furthermore, domain-based analysis was performed to assess its potential functional partners, revealing predicted interactions with ten proteins: Os02g0522700 protein (A0A0P0VJN4), Os05g0122400 protein (A0A0WHC1), Os05g0463900 protein (Q0DHI3_ORYSJ), Ubiquitin-40S ribosomal protein S27a-2 (RPS27AB), Ubiquitin-40S ribosomal protein S27a-1 (RPS27AA), Ubiquitin fusion protein (Q9FWV0_ORYSJ), Os02g0101700 protein (Q0E4T3_ORYSJ), Os09g0452700 protein (Q67V00_ ORYSJ), Os03g0259500 protein (A0A0P0VVP8), and Os07g0493033 protein (A0A0P0X666) (Fig. 4C).
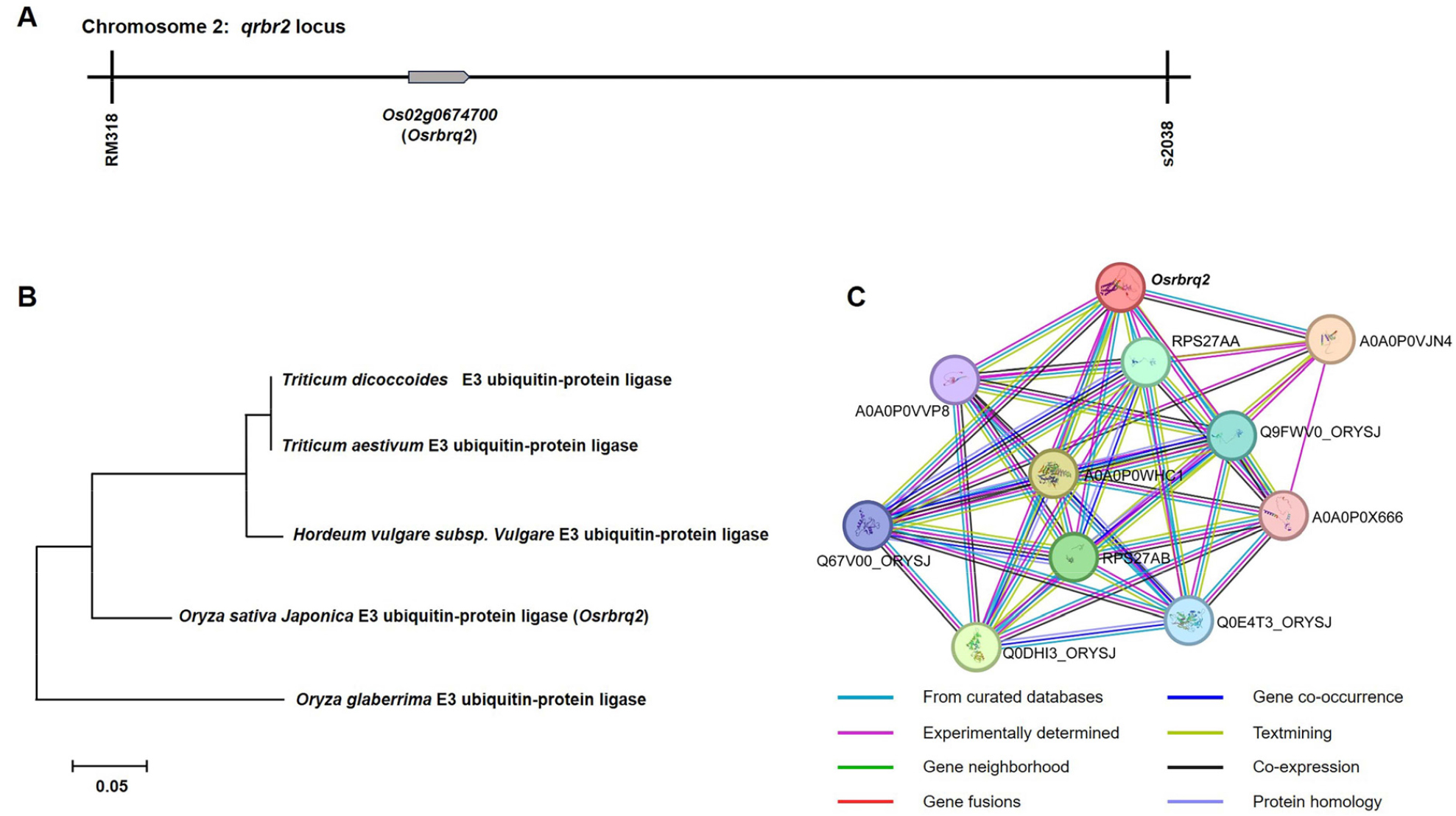
Fig. 4.
Phylogenetic and protein–protein interaction analysis of Osrbrq2. (A) The candidate gene Osrbrq2, identified as a target for rice blast resistance, was mapped within the qrbr2 locus, between the markers RM318-s2038 on chromosome 2. (B) Phylogenetic relationships of Osrbrq2 and its homologs were inferred using the parsimony method with 1,000 bootstrap replicates. (C) Protein–protein interaction analysis revealed that Osrbrq2 interacts with ten proteins: Os02g0522700 protein (A0A0P0VJN4), Os05g0122400 protein (A0A0WHC1), Os05g0463900 protein (Q0DHI3_ORYSJ), Ubiquitin-40S ribosomal protein S27a-2 (RPS27AB), Ubiquitin-40S ribosomal protein S27a-1 (RPS27AA), Ubiquitin fusion protein (Q9FWV0_ORYSJ), Os02g0101700 protein (Q0E4T3_ORYSJ), Os09g0452700 protein (Q67V00_ORYSJ), Os03g0259500 protein (A0A0P0VVP8), and Os07g0493033 protein (A0A0P0X666).
DISCUSSION
Rice blast remains a major constraint on global rice production, as the pathogen attacks leaves and panicles, leading to severe yield and quality losses (Ashkani et al., 2015). Although extensive breeding programs have incorporated resistance genes into elite cultivars, the rapid evolution and genetic variability of Magnaporthe oryzae frequently overcome single-gene resistance (Miah et al., 2013). The discovery of QTLs associated with blast resistance offers important insights into the genetic architecture of this trait and supports marker-assisted selection for more durable resistance. Nevertheless, the complexity of host–pathogen interactions indicates that deeper integration of QTL mapping with functional genomics will be crucial for elucidating the regulatory mechanisms governing resistance at both the leaf and panicle stages (Tian et al., 2022; Zhao et al., 2024).
In this study, we used rice blast isolates previously selected based on their pathogenic characteristics, including a set of 28 representative pathotypes identified from isolates collected in Korea between 2020 and 2022 (Zhao et al. 2026, in press). These isolates were used to inoculate the SNDH population, and QTL analysis was performed to identify genomic loci significantly associated with rice blast resistance. The SNDH population exhibited scores ranging from 1 to 5, reflecting responses from moderate resistance to high susceptibility. This clear phenotypic contrast between the parental lines confirms the suitability of the SNDH population for mapping rice blast-resistance loci. The observed variation among progeny provides a strong foundation for QTL identification, enabling the detection of both major and minor loci contributing to quantitative resistance in rice. In this study, QTL analysis revealed two loci linked to rice blast resistance, qrbr2 and qrbr6. Numerous mapping studies have identified blast-resistance loci across the rice genome, demonstrating that resistance is governed by major R genes and multiple QTLs with variable effects (Devanna et al., 2024; Wang et al., 2024; Zhang et al., 2021). Previous research has reported many QTLs associated with blast resistance distributed throughout the genome. For example, Jiang et al. (2020) identified 16 QTLs for blast resistance in the Jin23B/CR071 population, including qBR2–1 located between markers RM236 and RM451 on chromosome 2, and qBR6, positioned between RM539 and RM19951 on chromosome 6 (Jiang et al., 2020). A meta-QTL study by Devanna et al. (2024) identified several QTLs for blast resistance on chromosome 6, including MQTL6.3, which encompasses the Pi9/Piz gene cluster. This work further underscored the importance of chromosome 6 in conferring resistance to rice blast (Devanna et al., 2024). To date, more than 100 blast- resistance genes or QTLs have been reported, many of which are clustered on chromosomes 2, 6, 11, and 12, reflecting the evolutionary tendency of resistance genes to form tightly linked groups (Su et al., 2015; Vasudevan et al., 2015; Xiao et al., 2017; Zhao et al., 2024). In this context, our QTL analysis detected two significant novel loci, qrbr2 and qrbr6, which represent promising targets for future marker- assisted selection. GO analysis of the 774 genes located within these loci indicates that many candidate genes are associated with intracellular signaling, organelle organization, and cytoplasmic metabolic processes. These cellular functions are well known to contribute to plant defense responses, particularly through the regulation of reactive oxygen species homeostasis, signal transduction pathways, and pathogen recognition mechanisms (Waszczak et al., 2018).
Moreover, among the related genes identified within the loci qrbr2 and qrbr6, Os02g0674700 (Osrbrq2) was ultimately selected, as it encodes a RING-type E3 ubiquitin protein ligase—well-documented regulators of plant immune responses—and has been implicated in rice blast resistance. Although the specific function of Osrbrq2 has not yet been characterized, its annotation as a RING-type E3 ubiquitin ligase suggests a likely role in rice defense signaling. Previous studies have demonstrated that several RING-type E3 ligases, including APIP6, OsBBI1, and OsRGLG5, are essential for rice blast resistance (Park et al., 2012; Li et al., 2011; Liu et al., 2023), supporting our current findings and emphasizing the relevance of the qrbr2 and qrbr6 loci. Furthermore, protein–protein interaction analysis revealed that Osrbrq2 interacts with 10 other proteins; among these, Ubiquitin-40S ribosomal protein S27a-1 (RPS27AA) and Ubiquitin-40S ribosomal protein S27a-2 (RPS27AB), which encode ribosomal proteins of the S27 family, are known to play key roles in plant stress responses and translation regulation. Previous studies have shown that ribosomal proteins function not only in protein synthesis but also in stress adaptation and immune signaling pathways, underscoring their potential involvement in defense regulation (Fakih et al., 2023; Fakih & Germain, 2025). Thus, the Osrbrq2 gene identified in this study may contribute to rice blast resistance by modulating protein ubiquitination and by interacting with ribosomal proteins to regulate defense-related translation and stress signaling. Together, these functions indicate that Osrbrq2 represents a promising candidate gene associated with rice blast disease resistance.
However, further research is required to elucidate the molecular mechanisms underlying the role of Osrbrq2 in rice blast resistance. Functional characterization through CRISPR/Cas9- mediated knockout and transgenic overexpression, together with subcellular localization and effector-interaction assays, will be essential to validate its function and evaluate its potential for use in marker-assisted breeding aimed at achieving durable blast resistance. A deeper understanding of Osrbrq2-mediated defense signaling will not only clarify its contribution to host–pathogen interactions but also provide valuable genetic resources for developing rice cultivars with broad-spectrum and long-lasting resistance. Moreover, the detected loci qrbr2 and qrdr6 represent important genomic regions associated with rice blast resistance and offer promising targets for future functional characterization and breeding applications.
CONCLUSIONS
Rice blast remains one of the most destructive diseases affecting rice production, often causing substantial yield losses and posing a significant threat to global food security. In this study, we used a set of 28 representative isolates to inoculate the SNDH population for QTL analysis, which led to the identification of two major loci associated with rice blast resistance, qrbr2 and qrbr6. Within the qrbr2 locus, the candidate gene Os02g0674700 (Osrbrq2), encoding a RING-type E3 ubiquitin ligase, was identified as a promising resistance gene. The discovery of qrbr2,qrbr6, and Osrbrq2 provides valuable genetic resources for marker-assisted breeding and establishes a foundation for developing next-generation rice varieties with durable, long-term resilience to rice blast disease.